Bacteria growth in the chemostat.
| Download Bacteria3 desktop application (Windows 64bit) | Download Bacteria3 source code (Lazarus/Free Pascal) |
Description: Biology simulation, showing bacteria growth in the chemostat, based on the Monod growth model.
Simulation data is entered in the main window. Use Start to start the simulation: the number of bacteria pictures will increase/decrease
as time passes and the bar, showing the concentration of the substrate, indicates how this one is consumed (just see what happens if you increase the flow rate;
cf. included help for details). At application startup, parameters are set to update the simulation window each 0,1 sec, this simulation time corresponding to
1 min growth time in the real world chemostat; time update and interval may be changed in the Settings menu.
This visual part of the simulation is more programming fun of myself than really scientifically useful. If you are seriously interested in the simulation results,
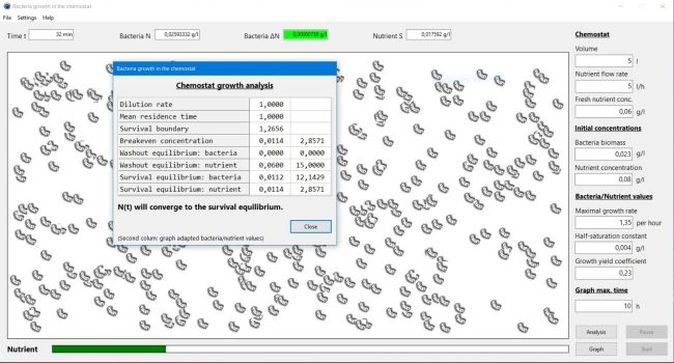
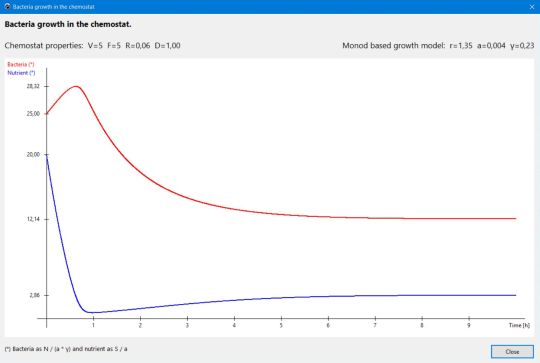
use the Graph button to display the variation of the bacteria biomass and the nutrients concentration vs. time. The Analyze button pops up a window, showing all important values (in particular the survival boundary; cf. help text) for the actual growth
simulation.
Free Pascal features: Creating images during program execution. Using a timer and the Left, Top and Visible property of images to create simple simulation programs. Using canvas to draw mathematical graphs.
Usage of string grids.
Screenshots:
|
|
If you like this application, please, support me and this website by signing my guestbook.