A general overview of the protein structure.
Please, note that the reader of the tutorial is supposed to have an intermediate knowledge of chemistry...
Proteins are the main macromolecules of an organism. When you look at an organism, what you see is either a protein or something that has been made by a protein. Structural proteins give the cell form and mobility, other proteins form pores in the cell membrane and control the traffic of small molecules into and out of the cell, and still other proteins regulate cellular activities in response to molecular signals from the external environment or from other cells. And all enzymes, i.e. biological catalysts that accelerate biochemical reactions in cells, are proteins.
1. Primary protein structure.
Proteins are polymers composed of monomers, called amino acids. In other words, a protein is a chain of amino acids. Because amino acids were once called peptides, the chain is also referred to as a polypeptide. All amino acids have an amino end (-NH2) and a carboxyl end (-COOH). They also have a side chain, called R (reactive) group. Amino acids general formula:
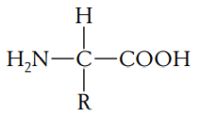 |
Of the many hundreds of described amino acids, 20 (22) are proteinogenic ("protein-building"). It is these 22 compounds that combine to give a vast array of peptides and proteins assembled by ribosomes (cf. Biological sequences and genetic code primer). Non-proteinogenic or modified amino acids may arise from post-translational modification or during nonribosomal peptide synthesis.
The carbon atom next to the carboxyl group is called the α–carbon. In proteinogenic amino acids, it bears the amine and the R group specific to each amino acid. With four distinct substituents, the α–carbon is stereogenic in all α-amino acids except glycine. All chiral proteinogenic amino acids have the L configuration.
The following picture (taken from the Wikipedia Amino acids article), shows the 21 proteinogenic α-amino acids found in eukaryotes, grouped according to their side chains.
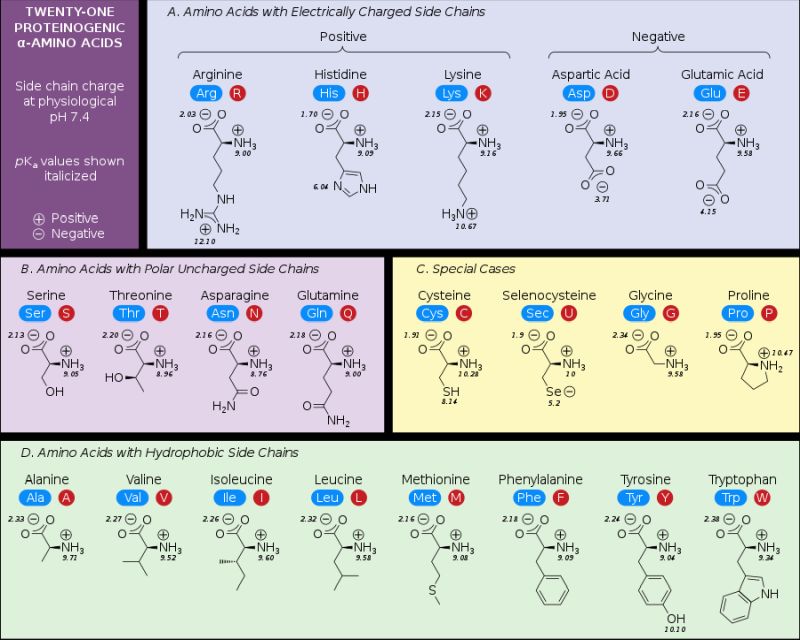 |
Amino acids sequences.
In proteins, the amino acids are linked together by covalent bonds called peptide bonds. A peptide bond is formed by the linkage of the -NH2 end of one amino acid with the -COOH end of another amino acid. Because of the way in which the peptide bond forms, a polypeptide chain always has an amino end and a carboxyl end.
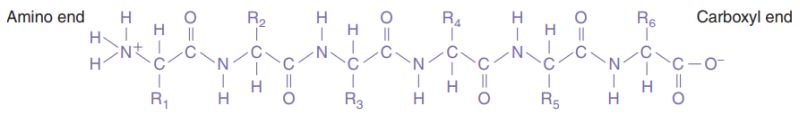 |
The primary structure (amino acid chain) is sufficient to uniquely identify it. This means that a protein may be described by a sequence of letters, where each letter codes for a given amino acid. Additional codes are X (to designate any/an unknown amino acid), B (for aspartic acid or asparagine), Z (for glutamic acid or glutamine), and J (leucine or isoleucine). These 1-letter codes are fine for being read by computer programs; to make a protein sequence better readable for humans, there are also 3-letter protein codes. Here is the complete table of standard IUB/IUPAC amino acid codes.
|
|
Note: Sec (U) and Pyl (O) are actually not part of the IUB/IUPAC amino acid codes, but selenocysteine and pyrrolysine are considered as standard amino acids by the Protein Database.
Amino acids classifications.
There are various ways to classify amino acids:
- By their molecule size: very small (Ala, Gly, Ser); small (Cys, Asp, Asn, Pro, Thr); medium (Glu, His, Gln, Val); large (Ile. Lys, Leu, Met, Arg); very large (Phe, Trp, Tyr).
- By their side chain structure: aliphatic (Ala, Ile, Gly, Leu, Val); aromatic (Phe, His, Trp, Tyr); cyclic (Pro), sulfur-containing (Cys, Met), acidic (Asp, Glu), basic (Lys, Arg), neutral (Asn, Gln, Ser, Thr).
- By their charge: positive (His, Lys, Arg); negative (Asp, Glu); uncharged (all others).
- By their polarity: polar (Asp, Glu, His, Lys, Asn, Gln, Arg, Ser, Thr, Tyr); nonpolar (Ala, Cys, Phe, Gly, Ile, Leu, Met, Pro, Val, Trp).
- By their hydropathy: hydrophobic (Ala, Cys, Phe, Ile, Leu, Met, Val, Trp); hydrophilic (Asp, Glu, Lys, Asn, Gln, Arg); neutral (Gly, His, Pro, Ser, Thr, Tyr).
- By their nutritive requirement (for humans): essential (Phe, Ile, Lys, Leu, Met, Thr, Val, Trp); semi-essential (His, Arg); non-essential (Ala, Cys, Asp, Glu, Gly, Asn, Pro, Gln, Ser, Tyr).
- By their chemical function: ketogenic (Lys, Leu, ); glucogenic & ketogenic (Phe, Ile, Thr, Trp, Tyr); glucogenic (all others).
Among the properties of amino acids, polarity and ionization determine their reactivity. Non-polar groups interact well with each other and poorly with polar groups. Positively charged amino acids are weak bases, fully protonated (Lys, Arg) or partly protonated (His) in normal biological conditions (pH = 7.0 - 7.4). Negatively charged amino acids are carboxylate groups, normally deprotonated at pH 7, and very polar.
Concerning the amino acids classifications, you might be interested in my Protein analysis: Amino acid count by similarity groups defined within the various classifications desktop application, shown on the screenshot below.
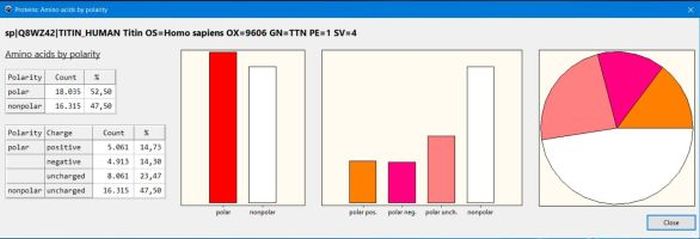 |
2. Secondary protein structure.
The secondary structure of a protein is the specific shape taken by the polypeptide chain by folding. In fact, the polypeptide backbone flexes by rotation about its single bonds. This results in regular repetitive structure arrangements, the most common being the α-helix and the β-sheet. The particular secondary structure is dependent on the local amino acid sequence. Amino acid can be grouped in three categories: amino acids with bulky side chains prefer to form a β-sheet, amino acids having side chains that disrupt the secondary structure, and amino acids tending to form an α-helix. Thus, we can define the secondary structure of a protein as the occurrence of regular repetitive patterns over short or longer sections of a polypeptide chain.
Flexing in a covalent chain structure may only be obtained by rotation about the axis of the C-C bonds. If we considered the polypeptide chain made only of single bonds (as suggests the picture in the "Amino acids sequences" section), this chain would be highly flexible, allowing all polypeptides to adopt a form that is random and disordered in bond orientation. Thus, the question: How to explain the existence of regular patterns?
Orderly arrangements of the polypeptide backbone were first studied by examining the fibrous proteins α-keratin and β-keratin (fibroin) using X-ray diffraction. X-ray diffraction showed the existence of a major and a minor pattern, each of them having a constant measurement (for a given of the two molecules). This led Linus Pauling to the hypothesis that the resonance structure of the peptide bond gives it double-bond characteristics. And he showed that considering these double bonds, the arrangement is normally a regular pattern. Normally, because it is also possible to set the peptide chain in a non-repetitive arrangement called a random coil. The picture below shows the double-bond characteristics of the peptide bond due to resonance. If you want to know more about this rather complex topic, have a look at BIOC*2580: Introduction to Biochemistry: Chapter 6 - Protein Secondary Structure at the Open Library website.
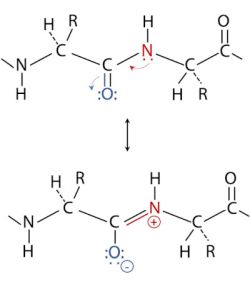 |
α-helices.
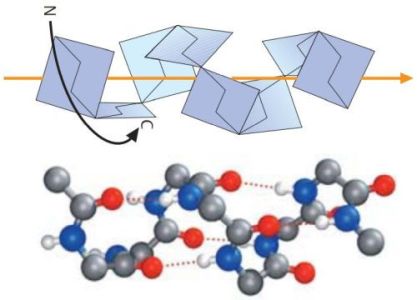
The right-handed α-helix is the most common type of secondary structure in proteins. In this conformation, the peptide chain is wound like a screw. Each turn of the screw covers approximately 3.6 amino acid residues, which means that there is one residue every 100 degrees of rotation (360/3.6). Each residue is translated 1.5 Å along the helix axis, which gives a vertical distance of 5.4 Å between structurally equivalent atoms in a turn (pitch of the screw). α-helices are stabilized by almost linear hydrogen bonds between the NH and CO groups of residues, which are four positions apart from each another in the sequence. In longer helices, most amino acid residues thus enter into two hydrogen bonds. On the picture1 below, oxygen atoms are represented in red, nitrogen atoms in blue, the hydrogen bonds as dotted orange lines; the full-color orange line represents the axis of the screw.
 |
Note: The mirror image of the α-helix, the left-handed α-helix, is rarely found in nature. Another type of helix occurs in the collagens. The left-handed collagen helix has a pitch of 0.96 nm and 3.3 residues per turn, and hydrogen bonds are not possible within this steeper arrangement. However, the conformation is stabilized by the association of three helices to form a right-handed collagen triple helix.
β-sheets.
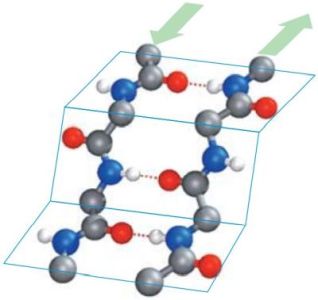
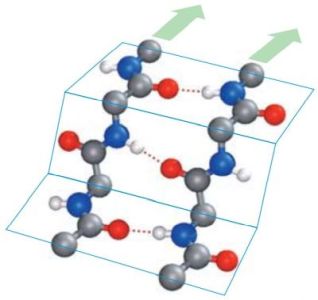
In β-pleated sheets, the peptide planes are arranged like a regularly folded sheet of paper. Again, hydrogen bonds can only form between neighboring chains. When the two strands run in opposite directions, the structure is referred to as an antiparallel pleated sheet (βa). When they run in the same direction, it is a parallel pleated sheet (βp). In both cases, the α-C atoms occupy the highest and lowest points in the structure, and the side chains point alternately straight up or straight down. Note that in extended pleated sheets, the individual chains of the sheet are usually not parallel, but twisted relative to one another. The pictures1 below, show an antiparallel β-sheet on the left, a parallel β-sheet on the right. Oxygen atoms are in red, nitrogen atoms in blue; the green arrows show the directions of the chains.
 |
 |
β-turns.
A β-turn is often found at a site where a peptide chain changes direction. These are sections in which four amino acid residues are arranged in such a way that the course of the chain reverses by about 180° into the opposite direction. β-turns types I and II are particularly frequent. Both are stabilized by hydrogen bonds between residues 1 and 4. β-turns are often located between the individual chains of antiparallel pleated sheets, or between chains of pleated sheets and α helices. The pictures1 below, show a type I β-turn on the left, a type II β-turn on the right. Oxygen atoms are in red, nitrogen atoms in blue.
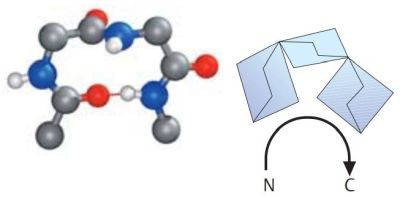 |
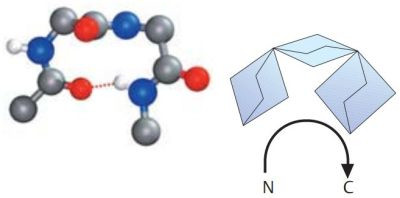 |
Other secondary structures.
Besides those mentioned above, there are several other secondary structures described, partially depending on given assignment categories. The most common are the π-helix and the 310-helix.
Secondary structure prediction.
To be done...
Supersecondary structure and protein motifs.
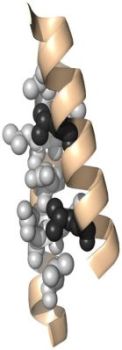
A supersecondary structure is often composed of two secondary structures linked together by a turn. These structures include: helix-turn-helix, helix-loop-helix, α-α corner, β-β corner, and β-hairpin. More complex supersecondary structures are: β-α-β motif, Zinc Finger motif, α-α motif (coiled coil), and others. The β-hairpin is common in proteins with β-sheet secondary structure that are formed of only two antiparallel chains, and results from a sharp hairpin turn (that does not disrupt the hydrogen bonding of the two β-pleated sheets). In contrast to the β-hairpin, a β-α-β motif contains two parallel β strands connected by a longer stretch of amino acids that form an α-helix. The Zinc Finger motif contains one zinc atom that stabilizes the fold by being centrally placed. In coiled coils, two α-helices are packed together by stably wounding around each other.
The pictures below show a β-hairpin structure (on the left), a helix-turn-helix structure (in the middle), and a α-α corner structure (on the right).

|
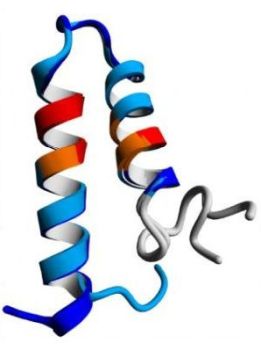
|

|
And here are pictures of a Zinc Finger motif (on the left), a β-α-β motif (in the middle), and a coiled coil motif (on the right).
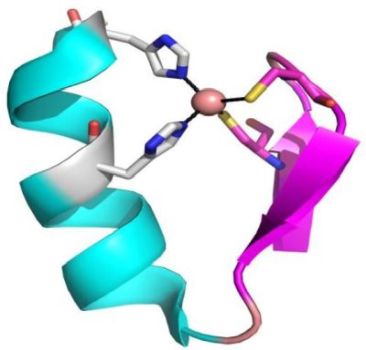
|

|
|
The term protein motifs, designating building blocks of protein structure is sometimes used as equivalent to the term supersecondary structures. Sometimes it refers to more complex repeated modalities visualized in many protein structures and created from secondary and supersecondary structural components. If you are interested in protein motifs and others details concerning protein structure, you might want to have a look at CH450 and CH451: Biochemistry - Chapter 2: Protein Structure at the website of Western Oregon University.
3. Tertiary protein structure.
Protein tertiary structure is the three-dimensional shape of a protein. The tertiary structure will have a single polypeptide chain backbone with one or more protein secondary structures. Amino acid side chains and the backbone may interact and bond in a number of ways. The interactions and bonds of side chains within a particular protein determine its tertiary structure.
A protein folded into its native state (native conformation) typically has a lower Gibbs free energy (a combination of enthalpy and entropy) than the unfolded conformation. Proteins will tend towards low-energy conformations, and this is what will determine the protein's fold in the cellular environment. Because many similar conformations have similar energies, protein structures are dynamic, fluctuating between these similar structures.
Fibrous proteins.
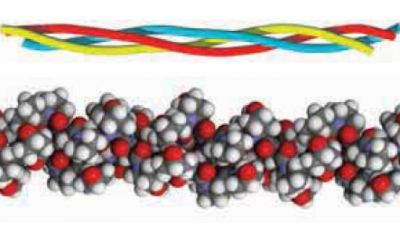
The simplest tertiary structure for a protein to adopt is a single uniform secondary structure. Examples: α-keratin is α-helix; fibroin (β-keratin) is an antiparallel β-sheet; collagen forms a unique triple helical structure, called the collagen helix. Secondary structures by themselves are rigid and give the protein an overall fibrous shape (whereas most proteins have a globular shape). The picture below shows the triple helical structure of the collagen molecule.
 |
Globular proteins.
A globular shape is the result of folding the protein onto itself. The "aim" of this folding is to bring the protein in its ideal folded state, i.e. the state where it is stabilized at a maximum. Here are some of the factors that "rule" protein folding:
- Clusters of amino acids that break the secondary structure of the protein (like Gly, Pro, Ser, Asn Asp) create turns (breaks of 2-3 residues, often having a well defined structure) and loops (longer section of amino acids with less regular arrangements), where the polypeptide can fold back on itself.
- The interactions within the tertiary structure is stabilized by the fact that non-polar or hydrophobic amino acids are grouped away from direct contact with water and occupy the core of the globular shape, whereas charged, hydrophilic residues, that interact well with the surrounding water, make up its surface. The grouping of non-polar amino acids together by hydrophobic interaction accounts for about 50% of the energy responsible for the stabilization of the folded form.
- The ideal folded state corresponds to an arrangement that maximizes the number of close atom to atom contacts. Atoms that are in perfect contact bind via a weak attractive force called the van der Waals interaction.
- Stabilizing interaction due to ion pairs: A negative charged side chain can pair up with a positive charged neighbor
- Stabilizing interaction due to H-bonds: These can form between donor groups (like Arg, Lys, His, Asn, Gln, Ser, Thr, Tyr) and acceptor groups (like Asp, Glu, Asn, Gln, His, Ser, Thr, Tyr).
- Stabilizing interaction due to disulfide bonds: These can form between pairs of Cys side chains that are aligned side by side.
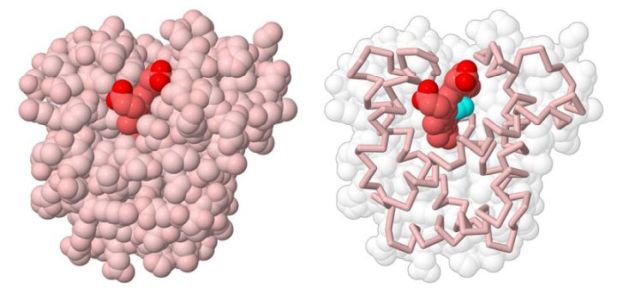
The picture below shows the globular tertiary structure of myoglobin.
 |
Protein domains.
Most proteins fold in a limited number of patterns, the protein structures usually arising from simple combinations of secondary structures (the supersecondary structures, that we saw in the previous section). These patterns are called protein domains. A domain is a compact three-dimensional structure, which often folds independently in a stable form such that it has its own hydrophobic core surrounded by hydrophilic residues. Domains form the fundamental unit of tertiary structure. Domain can evolve, function and exist independently. The size of a protein domain can vary in length from 30 to 500 amino acids.
During molecular evolution, domains are used as building blocks. These are recombined and assorted to form proteins with varied functions. A great variety of multidomain proteins can also be found which have arisen due to gene duplications. Independent proteins in prokaryotes can be seen as combination of domains in eukaryotic multidomain proteins.
Domains often have a specific function associated with them, thus large proteins can have multiple functional domains.
Several types of domains on the basis of secondary structures are:
- All α domains: Having only α-helices, they can be coiled coils, or helix bundles, composed of several α-helices held together by the hydrophobic core. Example: myoglobin.
- All β domains: Having only β-sheets, they can be divided into many groups. Example: superoxide dismutase (SOD).
- α + β domains: These domains contain a mixture of α and β secondary structures. Example: N-terminal domain of PAP phosphatase.
- α/β domains: These domains contain a combination of β-α-β motifs. It predominantly form parallel β-sheets surrounded by α-helices. Example: triosephosphate isomerase.
The pictures below show the superoxide dismutase (α + β domain; on the left) and the triosephosphate isomerase (α/β domains; on the right).
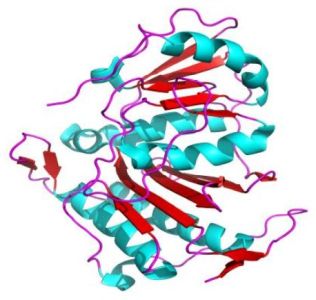 |
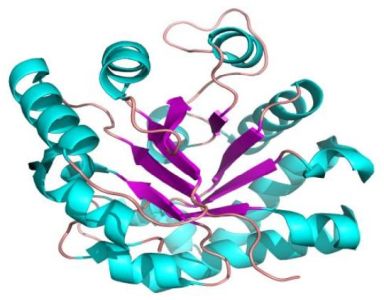 |
For further details concerning protein folding, stability and domains, you might want to have a look at BIOC*2580: Introduction to Biochemistry: Chapter 7 - Tertiary Structure & Protein Stability at the Open Library website.
Tertiary structure and protein function prediction.
To be done...
Quaternary protein structure.
Quaternary structure is the three-dimensional structure consisting of the aggregation of two or more individual polypeptide chains (subunits) that operate as a single functional unit. The resulting multimer is stabilized by the same non-covalent interactions and disulfide bonds as in tertiary structure. There are many possible quaternary structure organizations.
The picture shows a molecule of human hemoglobin, a tetramer of two α-globin and two β-globin subunits. The subunits are in red and blue, respectively, and the iron-containing heme groups in green.
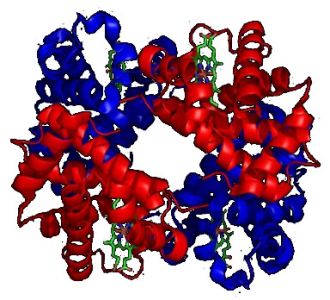 |
(1) Pictures taken from Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme.
If you find this tutorial helpful, please, support me and this website by signing my guestbook.